Research overview
My research program focuses on understanding drivers and processes of biodiversification. I'm especially interested in the way symbiotic interactions affect distribution, ecology and evolution of invertebrates.
Our lab mainly uses diverse mollusk groups (clams, cockles, snails, etc.) to address research questions, but other systems are also being explored (protists, algae, crustaceans, etc.). We adopt a combination of phylogenetic comparative methods, geometric morphometrics, molecular evolution and bioinformatics approaches to study biodiversification at species, population, and genomic levels.
Our lab mainly uses diverse mollusk groups (clams, cockles, snails, etc.) to address research questions, but other systems are also being explored (protists, algae, crustaceans, etc.). We adopt a combination of phylogenetic comparative methods, geometric morphometrics, molecular evolution and bioinformatics approaches to study biodiversification at species, population, and genomic levels.
Ongoing projects
|
Genomic evolution of photosymbiosis
Photosymbiotic associations between animal hosts and dinoflagellates are key components in maintaining the trophic and structural integrity of coral reef ecosystems. Although extensive efforts have been devoted to studying the ecology of such associations, the effects of photosymbiosis on marine metazoan genome evolution are largely unknown. The cockle subfamily Fraginae is an ideal group for addressing this topic. One lineage of Fraginae, the heart cockles and relatives, diverged from its non-symbiotic sister group around 30-40 million years ago and evolved symbiotic relationships with photosynthetic dinoflagellates. This allows a direct comparison of genomic compositions between symbiotic and non-symbiotic groups. Recent development of high-throughput sequencing technologies makes it possible to obtain large genomic/transcriptomic data from non-model organisms, which enables us to take advantage of statistical comparative methods and start to address the evolution of symbioses at the genomic level. Our lab uses comparative genomics approaches to identify candidate genomic adaptations to photosymbiosis in the Fraginae group and to explore their functions. Major questions being addressed are: 1. Which candidate genes are involved in the evolutionary adaptations to a symbiotic lifestyle in the symbiotic cockle lineage and their algal symbionts? 2. Which functional categories do those genes belong to and how have they been modified in the symbiotic species? The video on the right shows University of Guam student Megan Volsteadt collecting Fraginae bivalves in their typical habitats. |
|
|
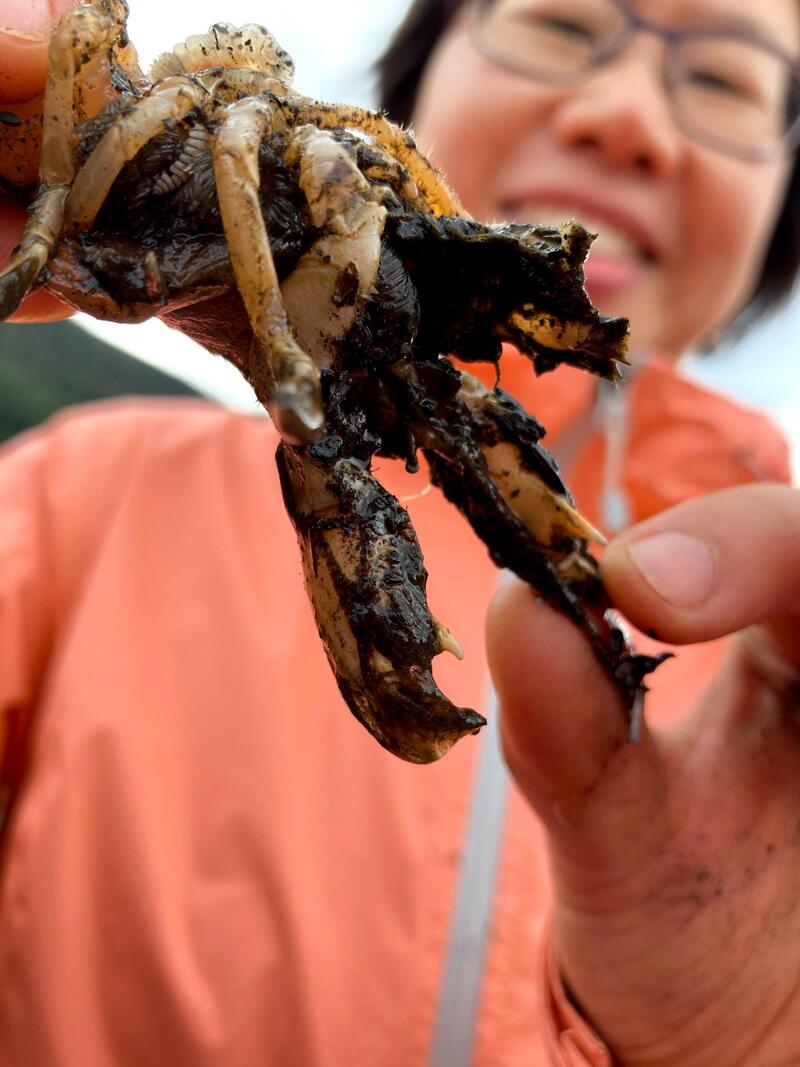
Invasive parasite ecology and host biogeography
In marine ecosystems, increased global-scale transportation creates opportunities for rapid introduction of invasive parasitic species. However, relevant research lags far-behind those focus on terrestrial and freshwater systems. This project will shed light on mechanisms that enable a successful marine parasite invasion. We will address three questions: (1) Do marine benthic microbiomes facilitate parasite-host recognition. (2) What makes introduced marine parasites more prevalent on newly-found hosts than their original hosts? (3) How do host populations maintain viable despite heavy parasite infection. Our study system is composed of the native blue mud shrimp (Upogebia pugettensis) from the Pacific coast of North America; a blood sucking isopod parasite (Orthione griffenis) introduced from Asia; and the original host Asian mud shrimp (Upogebia major), introduced separately to North America after the parasite has already established on the native shrimp. The native blue mud shrimp, Upogebia pugettensis, lives solitarily in permanent Y-shaped burrows on intertidal estuary mudflats, ranging from Alaska to California. The shrimps’ dense tube galleries form a unique and productive habitat on which dozens of native commensal species depend, including worms, crustaceans, clams, phoronids, and fish. U. pugettensis used to be spectacularly abundant and were classified as “ecosystem engineers” due to their critical function in nutrient cycling through sediment remineralization, large total biomass, and their great importance as prey for shorebirds and for juvenile salmon in estuaries. Most known U. pugettensis populations are declining or have collapsed following the invasion of Orthione griffenis in the early 1980s. O. griffenis live in the gill chambers of adult shrimp where they feed on shrimp hemolymph. Hemolymph losses effectively castrate up to 80% of infested females and causes populations to collapse. Losing Upogebia pugettensis, and the unique habitat that it creates, is as catastrophic to the eastern Pacific estuaries as the losses of tide flats, sea grass beds or marshes. Check out our fieldwork videos below. |
Mini documentary movies of our mud shrimp field trips
|
|
|
|
|
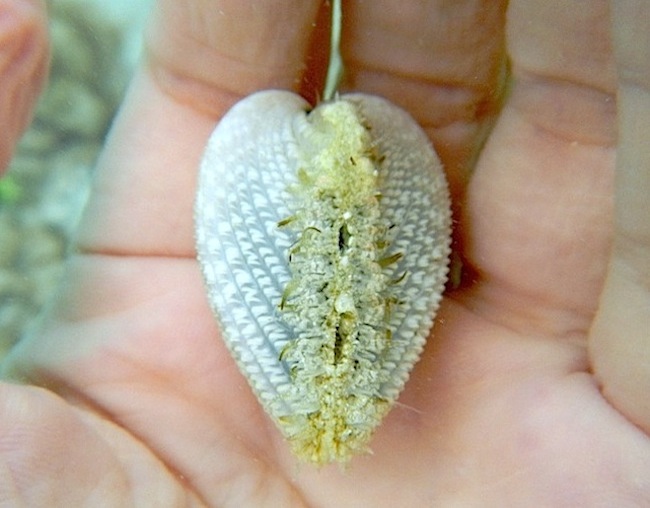
Macroevolution of symbiotic marine bivalves
How biotic interactions contribute to the generation and maintenance of global biodiversity is an emerging and vital question in biology. This project investigates the way symbiotic relationships shape long-term evolutionary dynamics of exceptionally diverse marine bivalve groups. The project focuses on the hyper-diverse marine bivalve superfamily Galeommatoidea. This bivalve group embodies a clear ecological dichotomy in that some members are free-living while others have obligate commensal associations with burrowing invertebrate hosts. Our lab generates molecular phylogenies of Galeommatoidea using hundreds of species. We use phylogenetic comparative approaches to compare rates of speciation and trait evolution between free-living and commensal groups. We also use this phylogenetic framework to test hypotheses regarding the role commensalism plays in the evolution other life history traits in Galeommatoidea. See a video on the left for an example of the bizarre commensal species. |
Others
Besides the above main projects, several collaborative projects are also going on in the lab, including phylogeography of freshwater snails in ancient lakes, conservation genetics of endangered island tree snails and other mollusks, physiology of symbiotic algae, and macroevolutionary process simulations. Learn more about them through my publications if you are interested.
Besides the above main projects, several collaborative projects are also going on in the lab, including phylogeography of freshwater snails in ancient lakes, conservation genetics of endangered island tree snails and other mollusks, physiology of symbiotic algae, and macroevolutionary process simulations. Learn more about them through my publications if you are interested.